Mitochondrial and endoplasmic reticulum redox biology
in health and diseas
Topics
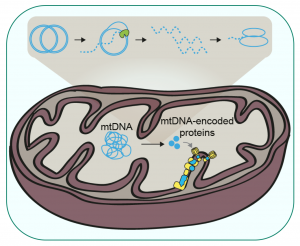
Regulation of mammalian mitochondrial gene expression
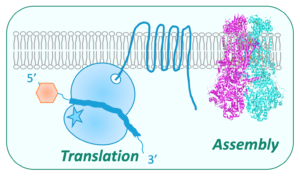
Post-translational steps of mitochondrial biogenesis in yeasts
PIs: Nathalie Bonnefoy and Geneviève Dujardin
The various steps of mitochondrial biogenesis including RNA metabolism, translation and OXPHOS complex assembly, are intimately linked and this coupling is facilitated by the fact that the mitochondrial ribosomes are bound to the inner-membrane. We mainly use budding and fission yeast models to dissect the molecular bases or investigate the cause of defects in these processes, identify new compensatory mechanisms, and search for drugs that could restore the respiratory function (Procaccio et al., 2019). We collaborate with I2BC platforms of high throughput sequencing (Naquin et al., 2018) and mass spectrometry (Lalève et al., 2020).
1-Mitochondrial translation represents a special focus since it is a key step of control in mitochondrial gene expression and it is affected in numerous human mitochondrial diseases. S. cerevisiae has been widely used for such studies and is a unique system since its mitochondria can be transformed to generate mitochondrial reporter genes. In addition, S. pombe is also a pertinent model because it (1) is highly dependent on respiration for survival, (2) contains the same set of general translation factors as humans, and (3) like human cells, has a compact mtDNA encoding mitochondrial mRNAs (mt-mRNAs) with limited untranslated regions. These similarities suggest that S. pombe and mammalian mt-mRNAs could be translated using very similar mechanisms. We have indeed found several examples of cross-complementation of mitochondrial translation mutants from both yeasts by human factors.
Our interest is focused on three classes of factors: (1) translation factors involved in the translation of all mt-mRNAs, more specifically termination and mitochondrial ribosome recycling (Ostojić et al., 2016), (2) subunits of the mitochondrial ribosome [current work], and (3) factors controlling the metabolism and/or translation of specific mt-mRNAs (Herbert et al., 2013). This includes PPR proteins (penta-tricopeptide repeats), which are generally RNA binding proteins specific to organelles like Ppr4 but also helicases and novel factors interacting with RNAs. A special emphasis is placed on genetic compensatory mechanisms and physical interactions, that both reveal functional networks (Ostojić et al., 2014).
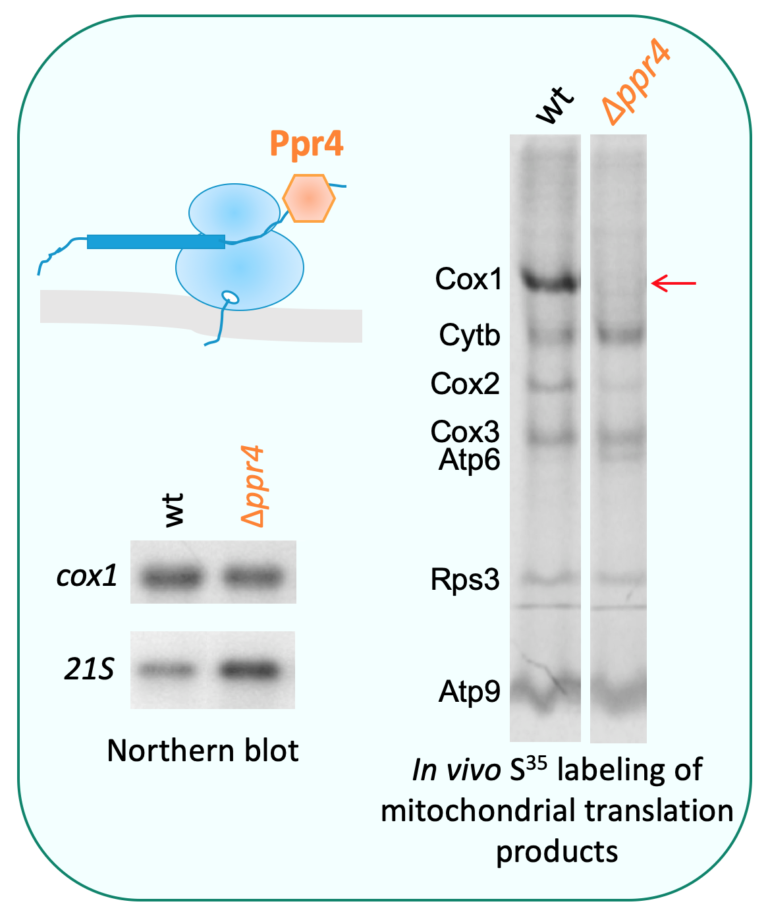
2-Assembly of the mitochondria- and nuclear-encoded subunits of the OXPHOS complexes is an intricate process involving numerous extrinsic assembly factors that execute highly specific functions.
For example, the chaperon Bcs1 (Panozzo et al., 2017) controls the assembly of the catalytic subunit Rip1 within the OXPHOS complex III. In addition, the respiratory complexes are organized into supramolecular structures or ‘‘super-complexes,’’ also called respirasomes, containing complexes I, III, and IV in higher eukaryotes and complexes III and IV in yeast.
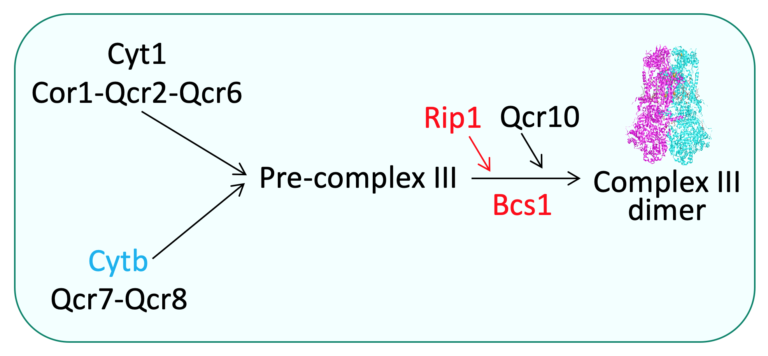
Most assembly factors are conserved through evolution. Mutations in the human genes encoding these assembly factors result in severe pathologies. We have introduced the corresponding mutations in the yeast genes encoding assembly factors of complex III or complex IV.
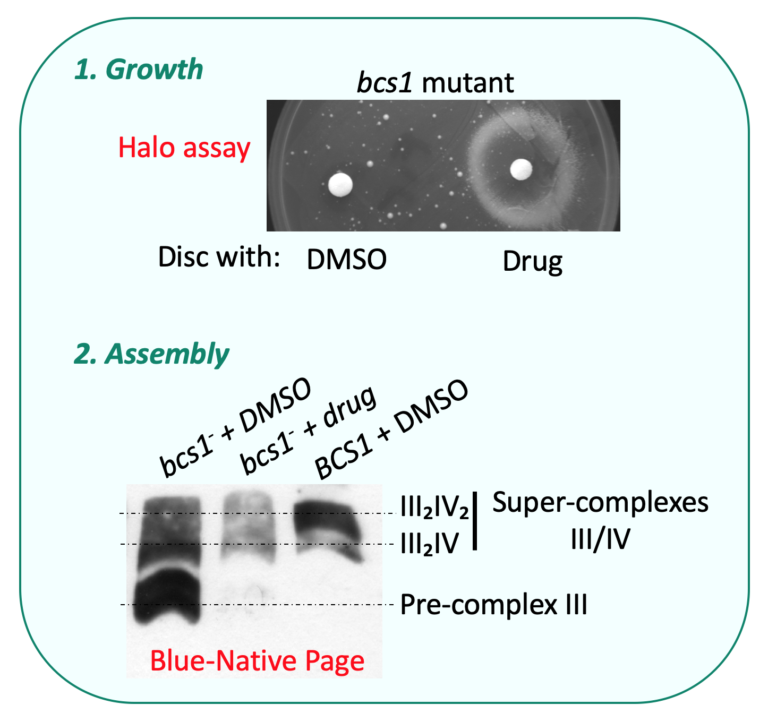
We have shown that mimicking some pathogenic mutations causes a respiratory deficiency that provides powerful means to screen for genetic or pharmacological compensations (Panozzo et al., 2017), that can be further tested in both yeast and mammals (Lasserre et al., 2015) (Lalève et al., 2020).
Our aim is to identify the targets of the compensatory molecules and compare them to genetic suppressors, in order to discover new actors or genetic interactions controlling the assembly of OXPHOS complexes.
Cited Publications
Herbert CJ, Golik P, Bonnefoy N. 2013. Yeast PPR proteins, watchdogs of mitochondrial gene expression. RNA Biol 10:1477–94. doi: 10.4161/rna.25392.
Lalève A, Panozzo C, Kühl I, Bourand-Plantefol A, Ostojic J, Sissoko A, Tribouillard-Tanvier D, Cornu D, Burg A, Meunier B, Blondel M, Clain J, et al. Bonnefoy N, Dujardin G. 2020. Artemisinin and its derivatives target mitochondrial c-type cytochromes in yeast and human cells. Biochim Biophys Acta – Mol Cell Res 1867:118661. doi: 10.1016/j.bbamcr.2020.118661.
Lasserre J-P, Dautant A, Aiyar RS, Kucharczyk R, Glatigny A, Tribouillard-Tanvier D, Rytka J, Blondel M, Skoczen N, Reynier P, Pitayu L, Rotig A, Delahodde A, Steinmetz LM, Dujardin G, Procaccio V, di Rago JP. 2015. Yeast as a system for modeling mitochondrial disease mechanisms and discovering therapies. Dis Model Mech 8:509–526. doi: 10.1242/dmm.020438.
Naquin D, Panozzo C, Dujardin G, VanDijk E, D’Aubenton-Carafa Y, C. T. 2018. Complete sequence of the intronless mitochondrial genome of the Saccharomyces cerevisiae strain CW252. Genome Announc 6:e00219-18. doi: doi.org/10.1128/genomeA .00219-18.
Ostojić J, Glatigny A, Herbert CJ, Dujardin G, Bonnefoy N. 2014. Does the study of genetic interactions help predict the function of mitochondrial proteins in Saccharomyces cerevisiae? Biochimie 100:27–37. doi: 10.1016/j.biochi.2013.11.004.
Ostojić J, Panozzo C, Bourand-Plantefol A, Herbert CJ, Dujardin G, Bonnefoy N. 2016. Ribosome recycling defects modify the balance between the synthesis and assembly of specific subunits of the oxidative phosphorylation complexes in yeast mitochondria. Nucleic Acids Res 44:5785–5797. doi: 10.1093/nar/gkw490.
Panozzo C, Laleve A, Tribouillard-Tanvier D, Ostojić J, Sellem CH, Friocourt G, Bourand-Plantefol A, Burg A, Delahodde A, Blondel M, Dujardin G. 2017. Chemicals or mutations that target mitochondrial translation can rescue the respiratory deficiency of yeast bcs1 mutants. Biochim Biophys Acta – Mol Cell Res 1864:2297–2307. doi: 10.1016/j.bbamcr.2017.09.003.
European patent from June 19th, 2019 n° 19305784.1, déposited by the AFM-Téléthon : « Use of Disulfiram or its derivatives for the treatment of mitochondrial diseases or dysfunction ». International extention through PCT: PCT/EP2020/067195. Inventors: Procaccio V, Rötig A, Delahodde A, Tribouillard-Tanvier D, Dujardin G, Blondel M; Organisms: AFM-téléthon, University & CHU Angers, INSERM, CNRS, Université de Bretagne Occidentale, AFM, Université de Paris, Université Paris-Saclay, Université de Bordeaux.
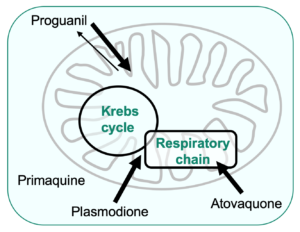
Mode of action of antimalarial compounds that target the mitochondria, using budding yeast as a model
PI: Brigitte Meunier
We are using the yeast model to investigate the mode of action of old and new antimalarial drugs that target the mitochondria, such as the biguanide proguanil that has been used for decades, primaquine, an 8-aminoquinoline developed in the 1940’s and the promising new lead plasmodione, a 3-benzyl-menadione, synthesized by the team of Elisabeth Davioud-Charvet (“Laboratoire d’Innovation Moléculaire et Applications”, University of Strasbourg).
Our approach is mainly genetics, combined with biochemical assays. The main objective is to identify primary targets of the drugs and to uncover candidate genes or pathways that would be involved in drug sensitivity/resistance.
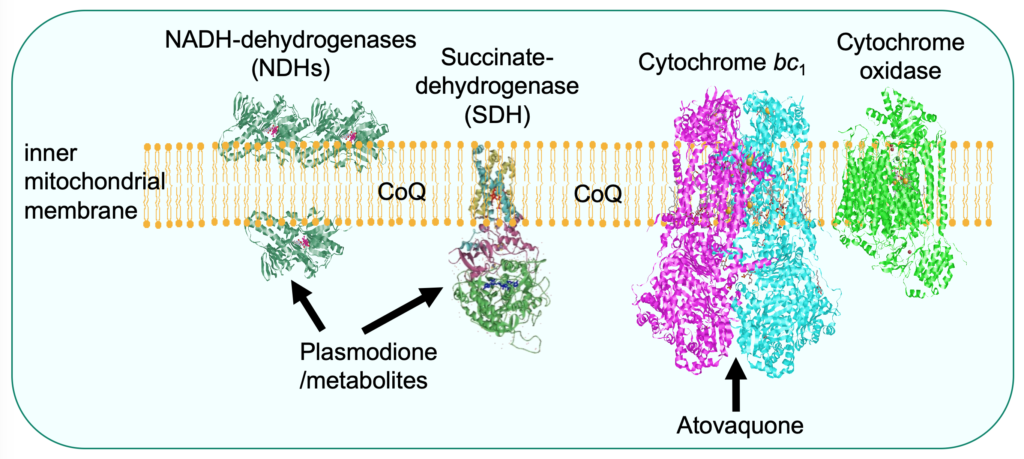
Proguanil is used in combination with its synergistic partner atovaquone, a well-known inhibitor of the mitochondrial respiratory chain cytochrome bc1 (Fisher et al., 2020). The mechanism of action of proguanil is still poorly understood. We proposed that the drug would not have a specific target but accumulate in the mitochondrial to concentrations that impair multiple mitochondrial functions leading to cell death. Analysis of proguanil resistant mutants led us to an unexpected resistance mechanism. We observed that modification of CoQ level resulted in increased or decreased proguanil sensitivity, by presumably, altering the mitochondrial membrane properties (Mounkoro et al., 2021).
Primaquine likely kills the parasites by a combination of redox-mediated mechanisms but the direct targets remain poorly characterized. The drug inhibits yeast respiratory growth and deletion of SOD2 encoding the mitochondrial superoxide dismutase severely increases its effect, which can be countered by the overexpression of AIM32 and MCR1 encoding mitochondrial enzymes involved in the response to oxidative stress. Further analysis suggested that ROS-labile Fe-S groups (as present in the Krebs cycle aconitase) would be targets of primaquine (Lalève et al., 2016).
Plasmodione is a redox-active compound that impairs the redox balance of parasites leading to cell death. A model of its mode of action was drawn, involving the generation of active metabolites that act as subversive substrates of flavoproteins, initiating a redox cycling process producing reactive oxygen species (Müller et al., 2011). The data obtained with the yeast model supports the proposal that plasmodione is a pro-drug acting via its benzhydrol and benzoyl metabolites. We showed that these metabolites act as substrate of the mitochondrial respiratory chain NADH-dehydrogenases that are, in yeast, the main redox-cycling enzymes (Mounkoro et al., 2019). Analysis of plasmodione resistant mutants then pointed towards another enzyme component of both the respiratory chain and the Krebs cycle, namely the succinate dehydrogenase (SDH). Based on the study of mutants, we proposed that the SDH-bound FAD co-factor would play a key role in plasmodione bioactivation, possibly in the first step of the process, highlighting a novel property of SDH (Mounkoro et al., 2020).
We are currently pursuing the project. To decipher further the drugs mode of action, we plan to broaden the search for resistant mutants and develop a strategy to uncover hypersensitive mutants. In parallel, more gene/protein candidates will also be tested. In addition, new compounds of the benzyl-menadione family have been synthesized in E. Davioud-Charvet’s lab, which we plan to investigate in yeast.
The findings obtained with yeast need be validated by study on the parasites. The data we obtained so far highlighted main metabolic pathways and enzymes that are well conserved between yeast and the parasite, stressing the relevance of the yeast model.
Cited Publications
Fisher N, Meunier B, Biagini GA. 2020. The cytochrome bc1 complex as an antipathogenic target. FEBS Lett 594:2935–2952. doi: 10.1002/1873-3468.13868
Lalève A, Vallières C, Golinelli-Cohen M-P, Bouton C, Song Z, Pawlik G, Tindall SM, Avery S V., Clain J, Meunier B. 2016. The antimalarial drug primaquine targets Fe–S cluster proteins and yeast respiratory growth. Redox Biol 7:21–29. doi: 10.1016/j.redox.2015.10.008.
Mounkoro P, Michel T, Blandin S, Golinelli-Cohen MP, Davioud-Charvet E, Meunier B. 2019. Investigating the mode of action of the redox-active antimalarial drug plasmodione using the yeast model. Free Radic Biol Med 141:269–278. doi: 10.1016/j.freeradbiomed.2019.06.026.
Mounkoro P, Michel T, Golinelli-Cohen MP, Blandin S, Davioud-Charvet E, Meunier B. 2020. A role for the succinate dehydrogenase in the mode of action of the redox-active antimalarial drug, plasmodione. Free Radic Biol Med. Epub ahead of print.
doi: 10.1016/j.freeradbiomed.2020.11.010.
Mounkoro P, Michel T, Meunier B. 2021. Revisiting the mode of action of the antimalarial proguanil using the yeast model. Biochem Biophys Res Commun 534:94–98. doi: 10.1016/j.bbrc.2020.12.004.
Müller T, Johann L, Jannack B, Brückner M, Lanfranchi DA, Bauer H, Sanchez C, Yardley V, Deregnaucourt C, Schrével J, Lanzer M, Schirmer RH, et al. 2011. Glutathione reductase-catalyzed cascade of redox reactions to bioactivate potent antimalarial 1,4-naphthoquinones – A new strategy to combat malarial parasites. J Am Chem Soc 133:11557–11571. doi: 10.1021/ja201729z.
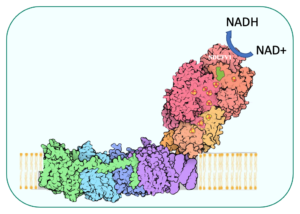
Complex I dysfunction and drug screening
PI: Carole Sellem
Complex I (nicotinamide adenine dinucleotide (NADH): ubiquinone oxidoreductase, EC 1.6.5.3) is the first and largest enzyme of the mitochondrial respiratory chain (RC) and oxidative phosphorylation (OXPHOS) system, that contains more than 45 subunits.
Isolated deficiency of complex I is the most commonly identified biochemical defect in childhood-onset mitochondrial disease, accounting for approximately a third of all cases of OXPHOS disorders.
At the moment there is no cure for complex I deficiency and options for treatment of complex I patients are only symptomatic.
The main objective of our work was to identify drugs able to restore complex I deficiencies. We are using the filamentous fungus Podospora anserina that contains a complex I similar to that of humans. We have constructed a mutant strain (Pa nuo-51) mimicking a human mutation in the first subunit of complex I (NDUFV1), responsible for a Leigh syndrome in human. We have used this thermosensitive P. anserina mutant to search for compounds able to alleviate its growth phenotype.
Screening a library of small FDA-approved molecules on the Pa nuo-51 mutant strain identified promising molecules.
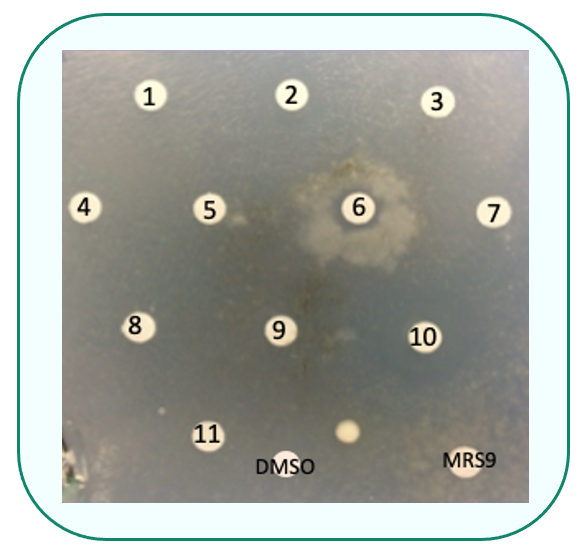

Drug drop test: Screening for molecules able to alleviate the mutant growth phenotype of Pa nuo-51at non permissive temperature
Two molecules (C1 and C2) were also validated in the corresponding mutant of C. elegans (nuo-1) and in cell cultures of patients (NDUFV1), since treatment increased respiration and complex I activity respectively.

Respiration and/or complex I activity in the mutant strain/cells of three model systems
The next step towards clinical assays is the validation of the molecules in NDUFV1 mouse models.
The identification of primary targets of the drugs is also a fundamental step that would help to improve our understanding of complex I regulation.
Ref:
SELLEM CH, Renaud A., Delahodde A., Procaccio V. (2020)
Use of Alverine or its derivative for the treatment of mitochondrial diseases or dysfunction associated with mitochondrial complex I deficiency.
Brevet déposé le 20/07/2020 à l’European Patent Office 80298 Munich Germany en 2020
Patent N°20305857.3 – 1112
