Interactions of Bacteria with Plants and Insects
Topics
Differentiation of endosymbiotic bacteria
Rhizobia are symbiotic bacteria able to infect leguminous plants and establish an intracellular mutualism in special root organs called nodules in which bacteria fix nitrogen and plants provide nutrients and protection. At the core of this symbiosis there is a bacterial differentiation program under the control of Nodule-specific Cysteine Rich peptides (NCR) which affects the bacterial physiology, in particular the metabolism, its cell cycle and DNA topology. The latter two aspects are objects of our investigation, using Sinorhizobium meliloti as symbiont of plants of the genus Medicago. In particular we investigate how the cell cycle regulatory machinery is involved in the differentiation and the role of genome topology in the bacteroid physiology.
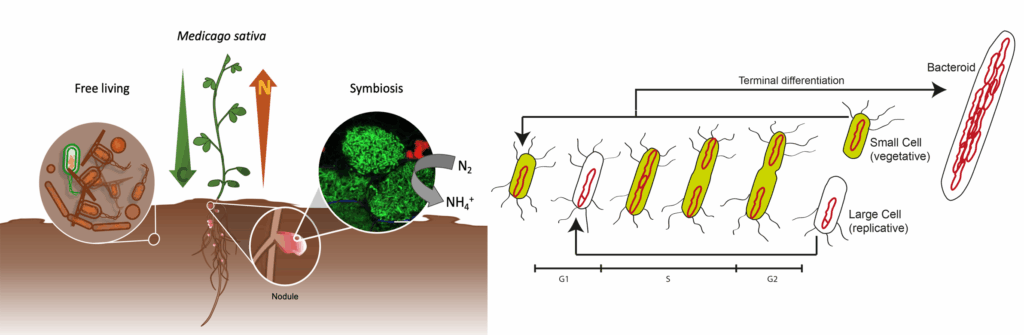
This project is financed by ANR SymGear, ANR Optileg, SPS CHroBaSym and I2BC Flagship program FungBugs
Selected publications
Dendene, Sara, Shuanghong Xue, Roza Mohammedi, et al. 2025. “Sinorhizobium meliloti FcrX Coordinates Cell Cycle and Division during Free-Living Growth and Symbiosis by a ClpXP-Dependent Mechanism.” Proceedings of the National Academy of Sciences of the United States of America 122 (11): e2412367122. https://doi.org/10.1073/pnas.2412367122.
diCenzo, George C., Lisa Cangioli, Quentin Nicoud, et al. 2022. “DNA Methylation in Ensifer Species during Free-Living Growth and during Nitrogen-Fixing Symbiosis with Medicago Spp.” mSystems 7(1): e01092-21. https://doi.org/10.1128/mSystems.01092-21.
Pini, Francesco, Benjamin Frage, Lorenzo Ferri, et al. 2013. “The DivJ, CbrA and PleC system controls DivK phosphorylation and symbiosis in Sinorhizobium meliloti.” Molecular Microbiology 90(1): 54-71. https://doi.org/10.1111/mmi.12347.
Pini, Francesco, Nicole J. De Nisco NJ, Lorenzo Ferri, et al. 2015. “Cell Cycle Control by the Master Regulator CtrA in Sinorhizobium meliloti.” PLoS Genetics 11(5):e1005232. https://doi.org/10.1371/journal.pgen.1005232.
Assembly of the gut microbiota in an insect model
The animal gut is the home of a microbiota composed of opportunistic members with a temporal presence, as well as highly specific species with a long-term presence, often located in a specific micro-niche of the gut. In a homeostatic interaction, the microbiota are usually beneficial for the host, providing services such as food digestion, nutrient provision, detoxcification of harmful compounds, immunity and even host behavior. We focus on the understanding of the role of the gut microbiota and of the host and bacterial mechanisms that underlie microbiota assembly and symbiont specificity in an insect model system, consisting of the bean bug Riptortus pedestris.


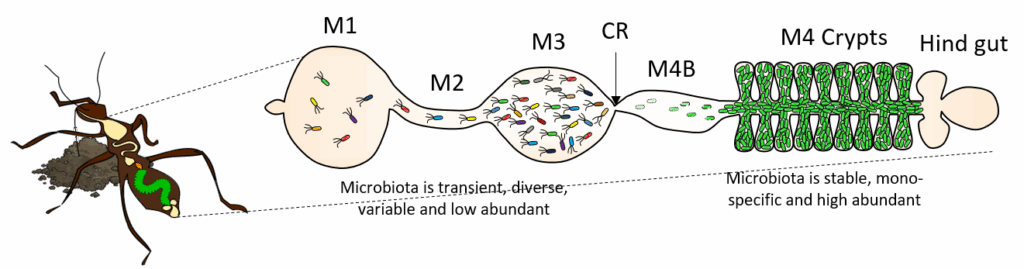
This project is funded by ANR, Graduate school Life Sciences and Health and OI MICROBES of the Paris-Saclay University, Saclay Plant Sciences and CNRS International Research Programme
Selected publications
Lextrait, Gaelle, Srotoswini Joardar, Raynald Cossard, Yoshitomo Kikuchi, Tsubasa Ohbayashi, and Peter Mergaert. 2025. “Strict Gut Symbiont Specificity in Coreoidea Insects Governed by Interspecies Competition within Caballeronia Strains.” The ISME Journal, November 13, wraf240. https://doi.org/10.1093/ismejo/wraf240.
Lachat, Joy, Gaëlle Lextrait, Romain Jouan, et al. 2024. “Hundreds of Antimicrobial Peptides Create a Selective Barrier for Insect Gut Symbionts.” Proceedings of the National Academy of Sciences 121 (25): e2401802121. https://doi.org/10.1073/pnas.2401802121.
Jang, Seonghan, Peter Mergaert, Tsubasa Ohbayashi, et al. 2021. “Dual Oxidase Enables Insect Gut Symbiosis by Mediating Respiratory Network Formation.” Proceedings of the National Academy of Sciences of the United States of America 118 (10). https://doi.org/10.1073/pnas.2020922118.
Kikuchi, Yoshitomo, Tsubasa Ohbayashi, Seonghan Jang, and Peter Mergaert. 2020. “Burkholderia Insecticola Triggers Midgut Closure in the Bean Bug Riptortus Pedestris to Prevent Secondary Bacterial Infections of Midgut Crypts.” The ISME Journal 14 (7): 1627–38. https://doi.org/10.1038/s41396-020-0633-3.
Ohbayashi, Tsubasa, Ryo Futahashi, Mia Terashima, et al. 2019. “Comparative Cytology, Physiology and Transcriptomics of Burkholderia Insecticola in Symbiosis with the Bean Bug Riptortus Pedestris and in Culture.” The ISME Journal 13 (6): 1469–83. https://doi.org/10.1038/s41396-019-0361-8.
The Agrobacterium tumefaciens lifestyle
The plant pathogen Agrobacterium tumefaciens is able to colonize the roots and galls (also named plant tumors) that it provokes on wounded host plants. While the virulence program of Agrobacterium tumefaciens has been deeply investigated during the past decades, little information is available about how A. tumefaciens exploits symptomatic and asymptomatic plant tissues in host and non-host plants. In this project, we combine global approaches (metabolomics, transcriptomics and Tn-Seq) to compare and decipher the Agrobacterium tumefaciens lifestyle in different ecological niches that are galls and roots of different plants, including Solanum lycopersicum and Arabidopsis thaliana.


This project is funded by ANR, OI Metabiodivex and Doctoral School SEVE of the Paris-Saclay University, SATT Paris-Saclay and I2BC Flagship program BacDrop
Selected publications
Torres, Marta, Audren Jiquel, Etienne Jeanne, Delphine Naquin, Yves Dessaux, and Denis Faure. 2021. “Agrobacterium Tumefaciens Fitness Genes Involved in the Colonization of Plant Tumors and Roots.” New Phytologist 233 (2): 905–18. https://doi.org/10.1111/nph.17810.
Faure, Denis. 2021. “Is There a Unique Integration Mechanism of Agrobacterium T-DNA into a Plant Genome?” The New Phytologist 229 (5): 2386–88. https://doi.org/10.1111/nph.17184.
Gonzalez-Mula, Almudena, Joy Lachat, Léo Mathias, et al. 2019. “The Biotroph Agrobacterium Tumefaciens Thrives in Tumors by Exploiting a Wide Spectrum of Plant Host Metabolites.” The New Phytologist 222 (1): 455–67. https://doi.org/10.1111/nph.15598.
González-Mula, Almudena, Julien Lang, Catherine Grandclément, et al. 2018. “Lifestyle of the Biotroph Agrobacterium Tumefaciens in the Ecological Niche Constructed on Its Host Plant.” The New Phytologist 219 (1): 350–62. https://doi.org/10.1111/nph.15164.
Plant protection and biocontrol
Since several years, we have undertaken a strong collaboration with the French professional organization of Seed Potato Growers (FN3PT) and its research branch (RD3PT) to improve potato protection strategies against the pectinolytic pathogens belonging to the genera Dickeya and Pectobacterium. Our collaborative researches are developing (1) knowledge on lifestyle and population dynamics of the pathogens, (2) molecular tools for pathogen diagnosis and detection and (3) biocontrol approaches.


This project is funded by ANR, Graduate school Life Sciences and Health and OI Living Machines à Work of the Paris-Saclay University, and I2BC Flagship program BacDrop.
Our team is hosting a full-time FN3PT engineer and successive PhDs and post-docs.
Partnership FN3PT/Inov3PT-CNRS 2017-2020 and 2021-2023
Selected publications
Robic, Kévin, Euphrasie Munier, Géraldine Effantin, et al. 2023. “Dissimilar Gene Repertoires of Dickeya Solani Involved in the Colonization of Lesions and Roots of Solanum Tuberosum.” Frontiers in Plant Science 14: 1154110. https://doi.org/10.3389/fpls.2023.1154110.
Blin, Pauline, Kévin Robic, Slimane Khayi, et al. 2022. “Pattern and Causes of the Establishment of the Invasive Bacterial Potato Pathogen Dickeya Solani and of the Maintenance of the Resident Pathogen D. Dianthicola.” Molecular Ecology 30 (2): 608–24. https://doi.org/https://doi.org/10.1111/mec.15751.
Oulghazi, Saïd, Jacques Pédron, Jérémy Cigna, et al. 2019. “Dickeya Undicola Sp. Nov., a Novel Species for Pectinolytic Isolates from Surface Waters in Europe and Asia.” International Journal of Systematic and Evolutionary Microbiology 69 (8): 2440–44. https://doi.org/10.1099/ijsem.0.003497.
Raoul des Essarts, Yannick, Jacques Pédron, Pauline Blin, Erwin Van Dijk, Denis Faure, and Frédérique Van Gijsegem. 2019. “Common and Distinctive Adaptive Traits Expressed in Dickeya Dianthicola and Dickeya Solani Pathogens When Exploiting Potato Plant Host.” Environmental Microbiology 21 (3): 1004–18. https://doi.org/10.1111/1462-2920.14519.
