Molecular Assemblies
and Genome Integrity
Topics
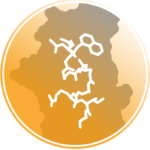
Maintenance of genome integrity relies on intricate cross-talks between DNA associated machineries, chromatin factors and signaling pathways regulated through versatile protein-protein interactions. Understanding the molecular logic underlying these interactions requires that competitive and synergistic associations be uncovered at different scales, from the atomic to the cellular one. We have been particularly interested in a class of proteins, called assembly chaperones which are involved in the formation of large macromolecular assemblies. As an example, we showed how Asf1, a central chaperone of histones H3 and H4, act in a synergistic manner with the Mcm2 helicase to coordinate the assembly and disassembly of nucleosomes at the replication fork. Other binding surfaces on Asf1 were also characterized revealing how other partners such as Rad53 (involved in signaling DNA damages), HirA (transcription) or CAF1 (replication) interact in a competitive manner with Asf1. These interactions take place in different epigenetic context and we are currently investigating how this context is influencing the network of interactions around Asf1.
Strong complementarity between experimental approaches and computational strategies drives the need for efficient structural prediction methods. There is a large gap bet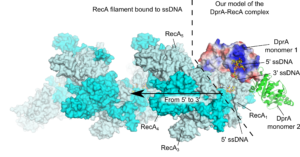 ween limited amounts of precise structural data and massive availability of sequence data. Using sequence information for structural prediction provides extremely powerful tools. In recent years, we have significantly contributed to improve methods for the prediction of protein complex structures by integrating coevolutionary information in the framework of the available docking tools.
ween limited amounts of precise structural data and massive availability of sequence data. Using sequence information for structural prediction provides extremely powerful tools. In recent years, we have significantly contributed to improve methods for the prediction of protein complex structures by integrating coevolutionary information in the framework of the available docking tools.
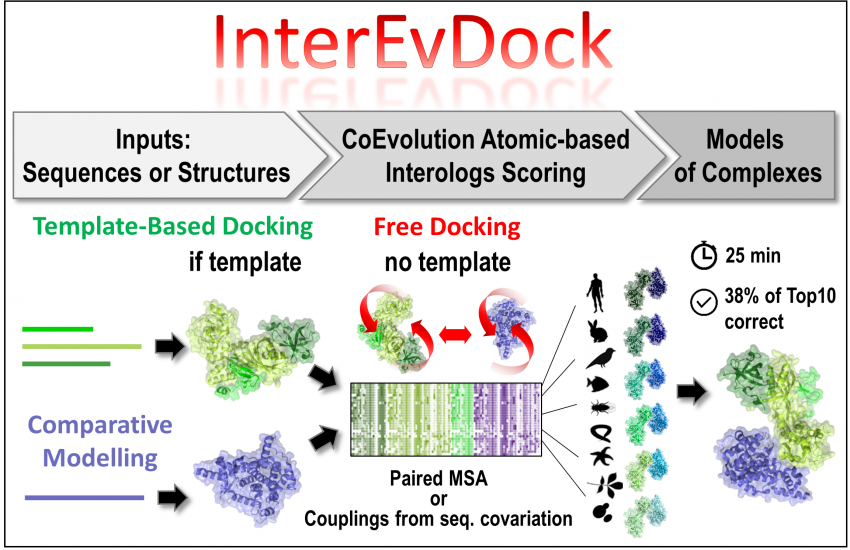
Through the development of the InterEvol database gathering a large number of complexes of known structure, we first performed an in-depth analysis of the co-evolution process at their interfaces under a structural perspective. In particular, we focused on more than 1000 pairs of homologous complexes, which allowed us to figure out important interface features at interfaces. From this analysis, we built the protein docking scoring function called InterEvScore by combining two and three-body statistical potentials with coevolutionary information. We further integrated this scoring function into a consensus of state-of-the-art prediction methods. Visit our server InterEvDock for a user-friendly interface to our latest developments in the field of docking. Thanks to these strategies, our team ranked among the top predictors of the 6th and 7th editions of the CAPRI (Critical Assessment of PRedicted Interactions) international challenges.
We are further trying to understand how we can improve the efficiency and precision of our docking methods through a better exploitation of the wealth of the information contained in multiple sequence alignments. We are also exploring integrative computational methods coupling omics data from high-throughput experiments with structural prediction of macromolecular complexes, including protein-RNA interactions. Our aim is to integrate structural models of interactions at the scale of modular assemblies and larger interaction networks.
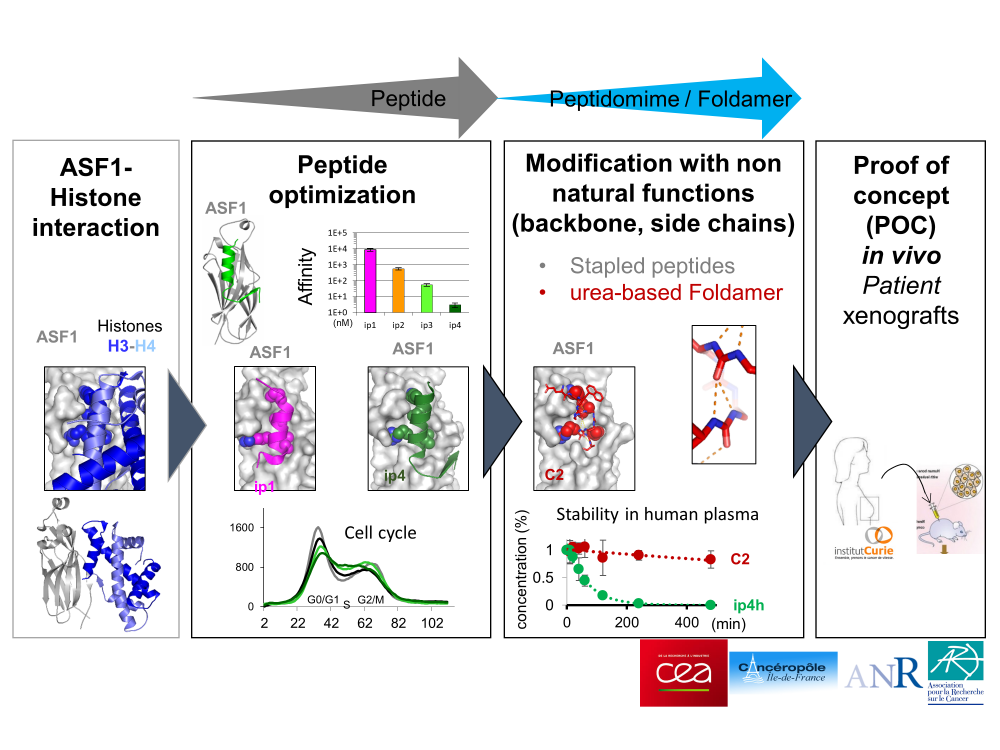
Design of protein-protein interaction inhibitiors
The inhibition of protein-protein interactions remains a significant methodological challenge which requires innovations in modeling, biomimetic chemistry, structural biology and cellular characterizations. Our team invested in this research field so as to inhibit the histone chaperone Asf1 and prevent its binding to its cognate partners the histones H3 and H4. Asf1 was recently recognized as a novel and promising anti-cancer target. We have developed a first generation of small peptides able to inhibit the interaction Asf1-histones. These peptides can penetrate cells efficiently, impede the cell cycle progression and trigger cell death in tumor cells cultures. Based on these encouraging results we are now tackling the conception of peptido-mimetic compounds, more powerful and selective, able to inhibit Asf1 activity in animals so as to demonstrate their therapeutic potential.